Tutorial about setting up an analysis pipeline and batch processing¶
Quite often you experiment with various analysis routines and appropriate parameters and come up with an analysis pipeline. A pipeline procedure then is a script defining analysis steps for a single locdata object (or a single group of corresponding locdatas as for instance used in 2-color measurements).
The Pipeline class can be used to combine the pipeline code, metadata and analysis results in a single pickleable object (meaning it can be serialized by the python pickle module).
This pipeline might then be applied to a number of similar datasets. A batch process is such a procedure for running a pipeline over multiple locdata objects and collecting and combing results.
from pathlib import Path
%matplotlib inline
import matplotlib.pyplot as plt
import locan as lc
lc.show_versions(system=False, dependencies=False, verbose=False)
Locan:
version: 0.22.0.dev32+g4bfc3ab8b
Python:
version: 3.11.14
# A path in which test data can be found:
TEST_DIR: Path = Path.cwd().parents[2] / "tests"
TEST_DIR
PosixPath('/home/docs/checkouts/readthedocs.org/user_builds/locan/checkouts/latest/tests')
Apply a pipeline of different analysis routines¶
Load rapidSTORM data file¶
path = TEST_DIR / 'test_data/npc_gp210.asdf'
print(path, '\n')
dat = lc.load_locdata(path=path, file_type=lc.FileType.ASDF)
/home/docs/checkouts/readthedocs.org/user_builds/locan/checkouts/latest/tests/test_data/npc_gp210.asdf
Jupyter environment detected. Enabling Open3D WebVisualizer.
[Open3D INFO] WebRTC GUI backend enabled.
[Open3D INFO] WebRTCWindowSystem: HTTP handshake server disabled.
dat.properties
{'localization_count': 202,
'position_x': np.float64(5623.488892810179),
'uncertainty_x': np.float64(3.9700846636030356),
'position_y': np.float64(6625.534602703435),
'uncertainty_y': np.float64(3.999303432482808),
'intensity': np.float32(3944778.0),
'local_background': np.float32(1131.3207),
'frame': np.int16(24),
'region_measure_bb': np.float32(47134.543),
'localization_density_bb': np.float32(0.004285604),
'subregion_measure_bb': np.float32(870.98047)}
Set up an analysis procedure¶
First define the analysis procedure (pipeline) in form of a computation function. Make sure the first parameter is the self refering to the Pipeline object. Add arbitrary keyword arguments thereafter. When finishing with return self the compute method can easily be called with instantiation.
def computation(self, locdata, n_localizations_min=4):
# import required modules
from locan.analysis import LocalizationPrecision
# prologue
self.file_indicator = locdata.meta.file.path
self.locdata = locdata
# check requirements
if len(locdata)<=n_localizations_min:
return None
# compute localization precision
self.lp = LocalizationPrecision().compute(self.locdata)
return self
Run the analysis procedure¶
Instantiate a Pipeline object and run compute():
pipe = lc.Pipeline(computation=computation, locdata=dat, n_localizations_min=4).compute()
pipe.meta
Processed frames:: 0%| | 0/24884 [00:00<?, ?it/s]
Processed frames:: 9%|▉ | 2315/24884 [00:00<00:00, 22664.82it/s]
Processed frames:: 25%|██▌ | 6245/24884 [00:00<00:00, 31941.51it/s]
Processed frames:: 46%|████▋ | 11539/24884 [00:00<00:00, 41067.36it/s]
Processed frames:: 100%|██████████| 24884/24884 [00:00<00:00, 62685.96it/s]
identifier: "1"
method {
name: "Pipeline"
parameter: "{\'computation\': <function computation at 0x75363eca2de0>, \'locdata\': <locan.data.locdata.LocData object at 0x753640b05490>, \'n_localizations_min\': 4}"
}
creation_time {
seconds: 1777538138
nanos: 593531000
}
Results are available from Pipeline object in form of attributes defined in the compute function:
[attr for attr in dir(pipe) if not attr.startswith('__') and not attr.endswith('__')]
['_get_parameters',
'_init_meta',
'_update_meta',
'computation',
'computation_as_string',
'compute',
'count',
'file_indicator',
'kwargs',
'locdata',
'lp',
'meta',
'parameter',
'report',
'results',
'save_computation']
pipe.lp.results.head()
| position_delta_x | position_delta_y | position_distance | original_index | frame | |
|---|---|---|---|---|---|
| 0 | -11.189941 | 4.859863 | 12.199716 | 0 | 24 |
| 1 | 32.580078 | 4.170410 | 32.845909 | 1 | 25 |
| 2 | 13.549805 | -15.439941 | 20.542370 | 3 | 141 |
| 3 | 4.669922 | 3.010254 | 5.556060 | 4 | 142 |
| 4 | 20.469727 | 14.750000 | 25.230383 | 6 | 239 |
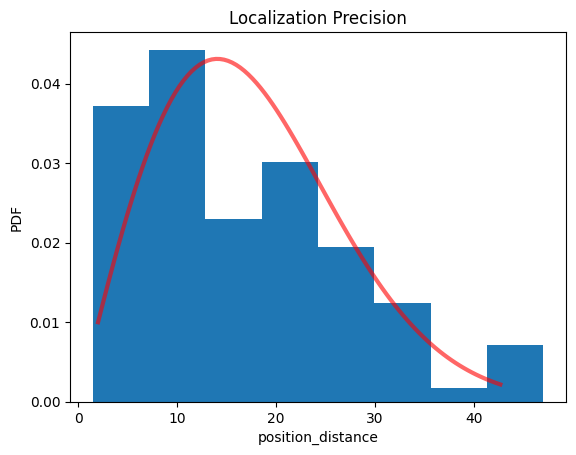
pipe.lp.hist();
print(pipe.lp.distribution_statistics.parameter_dict())
{'position_delta_x_loc': np.float32(0.6029139), 'position_delta_x_scale': np.float32(13.2682), 'position_delta_y_loc': np.float32(-1.227687), 'position_delta_y_scale': np.float32(14.76087), 'position_distance_sigma': np.float64(14.067675781250028), 'position_distance_loc': 0, 'position_distance_scale': 1}
You can recover the computation procedure:
pipe.computation_as_string()
'def computation(self, locdata, n_localizations_min=4):\n \n # import required modules\n from locan.analysis import LocalizationPrecision\n \n # prologue\n self.file_indicator = locdata.meta.file.path\n self.locdata = locdata\n \n # check requirements\n if len(locdata)<=n_localizations_min:\n return None\n \n # compute localization precision\n self.lp = LocalizationPrecision().compute(self.locdata)\n \n return self\n'
or save it as text protocol:
The Pipeline object is pickleable and can thus be saved for revisits.
Apply the pipeline on multiple datasets - a batch process¶
Let’s create multiple datasets:
path = TEST_DIR / 'test_data/npc_gp210.asdf'
print(path, '\n')
dat = lc.load_locdata(path=path, file_type=lc.FileType.ASDF)
locdatas = [lc.select_by_condition(dat, f'{min}<index<{max}') for min, max in ((0,100), (101,202))]
locdatas
/home/docs/checkouts/readthedocs.org/user_builds/locan/checkouts/latest/tests/test_data/npc_gp210.asdf
[<locan.data.locdata.LocData at 0x75369416a250>,
<locan.data.locdata.LocData at 0x753637ae0b10>]
Run the analysis pipeline as batch process
pipes = [lc.Pipeline(computation=computation, locdata=dat).compute() for dat in locdatas]
Processed frames:: 0%| | 0/6164 [00:00<?, ?it/s]
Processed frames:: 24%|██▍ | 1492/6164 [00:00<00:00, 14652.04it/s]
Processed frames:: 51%|█████▏ | 3166/6164 [00:00<00:00, 15754.07it/s]
Processed frames:: 77%|███████▋ | 4742/6164 [00:00<00:00, 13678.53it/s]
Processed frames:: 100%|██████████| 6164/6164 [00:00<00:00, 15603.97it/s]
Processed frames:: 0%| | 0/18639 [00:00<?, ?it/s]
Processed frames:: 5%|▌ | 1022/18639 [00:00<00:01, 10056.16it/s]
Processed frames:: 25%|██▍ | 4569/18639 [00:00<00:00, 24841.05it/s]
Processed frames:: 56%|█████▋ | 10511/18639 [00:00<00:00, 40173.02it/s]
Processed frames:: 97%|█████████▋| 18132/18639 [00:00<00:00, 53446.85it/s]
Processed frames:: 100%|██████████| 18639/18639 [00:00<00:00, 43773.67it/s]
As long as the batch procedure runs in a single computer process, the identifier increases with every instantiation.
[pipe.meta.identifier for pipe in pipes]
['2', '3']
Visualize the combined results¶
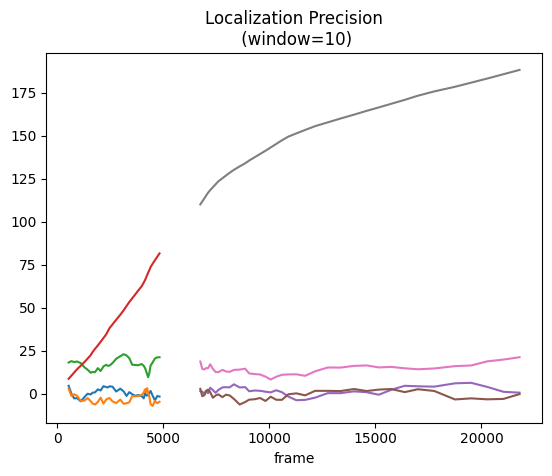
fig, ax = plt.subplots(nrows=1, ncols=1)
for pipe in pipes:
pipe.lp.plot(ax=ax, window=10)
plt.show()