Tutorial about analyzing blink statistics¶
SMLM depends critically on the fluorescence intermittency or, in other words, the blinking of fluorescence dyes. To characterize blinking properties you can compute on- and off-periods from clustered localizations assuming that they originate from the same fluorophore.
from pathlib import Path
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import scipy.stats as stats
import locan as lc
lc.show_versions(system=False, dependencies=False, verbose=False)
Locan:
version: 0.22.0.dev32+g4bfc3ab8b
Python:
version: 3.11.14
Synthetic data¶
We use synthetic data that represents localizations from a single fluorophore being normal distributed in space and emitting at a constant intensity. We assume that the on- and off-times in units of frames are distributed like a geometric distribution with a mean on_period mean_on and a mean off_period mean_off. Typically a geometric distribution is parameterized by a variable p with p = 1 / mean.
rng = np.random.default_rng(seed=1)
n_samples = 10_000
mean_on = 5
mean_off = 20
on_periods = stats.geom.rvs(p=1/mean_on, size=n_samples, random_state=rng)
off_periods = stats.geom.rvs(p=1/mean_off, size=n_samples, random_state=rng)
On- and off-times are converted in a series of frame numbers at which a localization was detected.
def periods_to_frames(on_periods, off_periods):
"""
Convert on- and off-periods into a series of increasing frame values.
"""
on_frames = np.arange(np.sum(on_periods))
cumsums = np.r_[0, np.cumsum(off_periods)[:-1]]
add_on = np.repeat(cumsums, on_periods)
frames = on_frames + add_on
return frames[:len(on_periods)]
frames = periods_to_frames(on_periods, off_periods)
offspring = [rng.normal(loc=0, scale=10, size=(n_samples, 2))]
locdata = lc.simulate_cluster(centers=[(50, 50)], region=[(0, 100), (0, 100)], offspring=offspring, clip=False, shuffle=False, seed=rng)
locdata.dataframe['intensity'] = 1
locdata.dataframe['frame'] = frames
locdata = lc.LocData.from_dataframe(dataframe=locdata.data)
print('Data head:')
print(locdata.data.head(), '\n')
print('Summary:')
locdata.print_summary()
print('Properties:')
print(locdata.properties)
Jupyter environment detected. Enabling Open3D WebVisualizer.
[Open3D INFO] WebRTC GUI backend enabled.
[Open3D INFO] WebRTCWindowSystem: HTTP handshake server disabled.
Data head:
position_x position_y cluster_label intensity frame
0 50.507149 36.698297 0 1 0
1 44.539062 27.969248 0 1 1
2 56.876954 42.152826 0 1 2
3 38.878363 43.999842 0 1 3
4 61.001644 54.231150 0 1 4
Summary:
identifier: "2"
comment: ""
source: DESIGN
state: RAW
element_count: 10000
frame_count: 10000
creation_time {
2026-04-30T08:33:17.171062Z
}
Properties:
{'localization_count': 10000, 'position_x': np.float64(49.986893963647944), 'uncertainty_x': np.float64(0.09881426457266232), 'position_y': np.float64(49.84659129517861), 'uncertainty_y': np.float64(0.10007238629130723), 'intensity': np.int64(10000), 'frame': np.int64(0), 'region_measure_bb': np.float64(6466.041774468444), 'localization_density_bb': np.float64(1.5465411992056104), 'subregion_measure_bb': np.float64(322.0177777257221)}
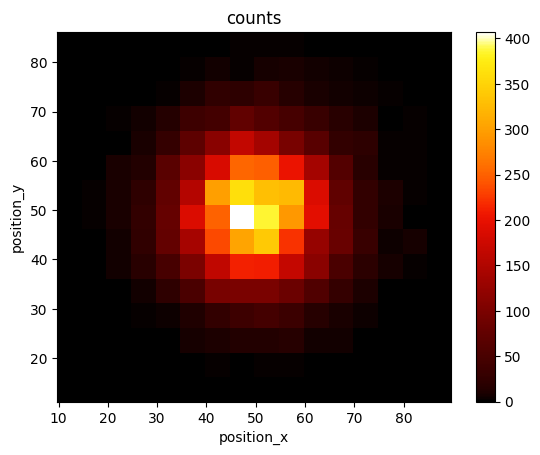
lc.render_2d(locdata, bin_size=5);
Blinking statistics¶
To determine on- and off-times for the observed blink events use the analysis class BlinkStatistics.
bs = lc.BlinkStatistics(memory=0, remove_heading_off_periods=False).compute(locdata)
bs
BlinkStatistics(memory=0, remove_heading_off_periods=False)
bs.results.keys()
dict_keys(['on_periods', 'on_periods_frame', 'off_periods', 'off_periods_frame', 'on_periods_indices'])
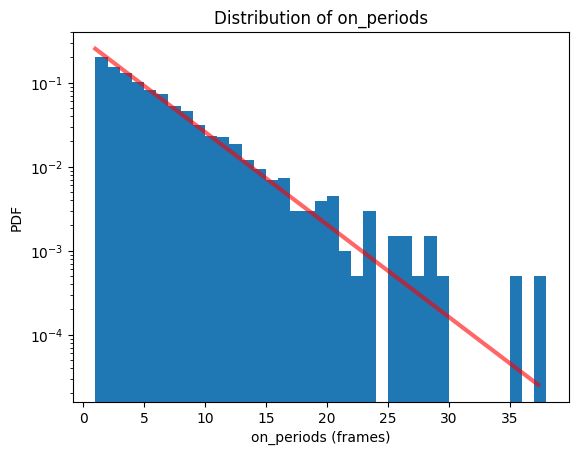
When plotting the histogram an exponential distribution is fitted by default.
bs.hist(data_identifier='on_periods');
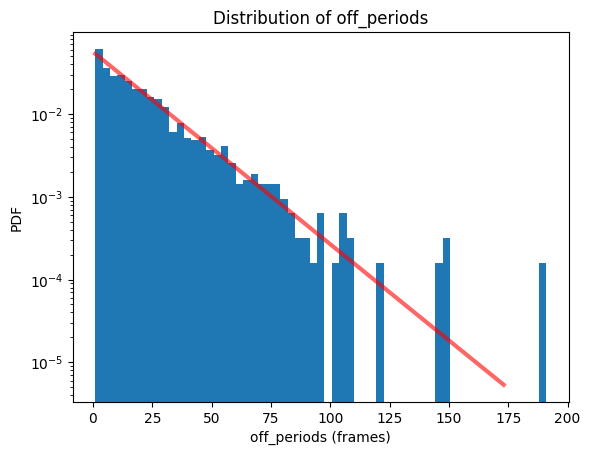
bs.hist(data_identifier='off_periods');
bs.distribution_statistics
{'on_periods': _DistributionFits(analysis_class=BlinkStatistics, distribution=expon_gen, data_identifier=on_periods),
'off_periods': _DistributionFits(analysis_class=BlinkStatistics, distribution=expon_gen, data_identifier=off_periods)}
The fit results provide loc and scale parameter (see scipy.stats documentation). For loc = 0, scale describes the mean of the distribution..
bs.distribution_statistics['on_periods'].parameter_dict()
{'on_periods_loc': 1.0, 'on_periods_scale': 3.9455984174085064}
bs.distribution_statistics['off_periods'].parameter_dict()
{'off_periods_loc': 1.0, 'off_periods_scale': 18.682830282038594}
Due to the default setting for the scaling parameter loc the mean on_period is on_periods_scale + on_periods_loc in agreement with our input value.
Geometric distribution¶
We can compare this with a geometric distribution that is estimated from the observed mean on_period on_periods_mean.
on_periods_mean = bs.results['on_periods'].mean()
on_periods_mean.round(2)
np.float64(4.95)
off_periods_mean = bs.results['off_periods'].mean()
off_periods_mean.round(2)
np.float64(19.68)
# test result
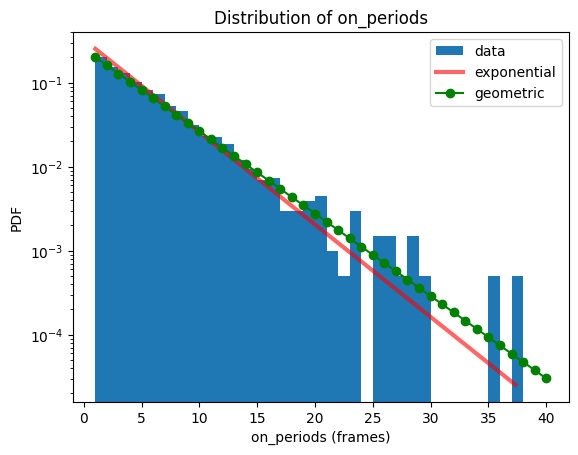
x = np.arange(stats.geom.ppf(0.01, 1/on_periods_mean), stats.geom.ppf(0.9999, 1/on_periods_mean))
y = stats.geom.pmf(x, 1/on_periods_mean)
fig, ax = plt.subplots()
bs.hist(data_identifier='on_periods', fit=False, label='data')
bs.distribution_statistics['on_periods'].plot(label='exponential')
ax.plot(x, y, '-go', label='geometric')
ax.set_yscale('log')
ax.legend(loc='best')
plt.show()